75
Transcriptional Regulation
Gene expression can be regulated at the transcriptional level. This means that the process of transcription can be turned on or off. In both prokaryotic and eukaryotic cells, transcription requires RNA polymerase to bind to a sequence upstream of a gene to initiate transcription. Prokaryotes almost always regulate gene expression at the transcriptional level.
In eukaryotes, the eukaryotic RNA polymerase requires other proteins, or transcription factors, to facilitate transcription initiation. Transcription factors are proteins that bind to the promoter sequence and other regulatory sequences to control the transcription of the target gene. RNA polymerase by itself cannot initiate transcription in eukaryotic cells. Transcription factors must bind to the promoter region first and recruit RNA polymerase to the site for transcription to be established.
- If transcription factors are not allowed to bind, transcription can not take place, which means gene expression is turned off.
- In some eukaryotic genes, there are regions that help increase or enhance transcription called enhancers. When transcription factors bind to these enhancer regions, they can increase rates of transcription, which means gene expression is turned up.
- Transcriptional repressors can bind to promoter or enhancer regions and block transcription. This means that gene expression is turned off.
Post-transcriptional Regulation
After RNA is transcribed, it must be processed into a mature form before translation can begin. This processing after an RNA molecule has been transcribed, but before it is translated into a protein, is called post-transcriptional modification. As with the epigenetic and transcriptional stages of processing, this post-transcriptional step can also be regulated to control gene expression in the cell. If the RNA is not processed, shuttled, or translated, then no protein will be synthesized.
Alternative RNA Splicing
In the 1970s, genes were first observed that exhibited alternative RNA splicing. Alternative RNA splicing is a mechanism that allows different protein products to be produced from one gene when different combinations of introns (and sometimes exons) are removed from the transcript (Figure 1). This alternative splicing can be haphazard, but more often it is controlled and acts as a mechanism of gene regulation, with the frequency of different splicing alternatives controlled by the cell as a way to control the production of different protein products in different cells, or at different stages of development. Alternative splicing is now understood to be a common mechanism of gene regulation in eukaryotes; according to one estimate, 70% of genes in humans are expressed as multiple proteins through alternative splicing.
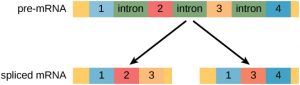
How could alternative splicing evolve? Introns have a beginning and ending recognition sequence, and it is easy to imagine the failure of the splicing mechanism to identify the end of an intron and find the end of the next intron, thus removing two introns and the intervening exon. In fact, there are mechanisms in place to prevent such exon skipping, but mutations are likely to lead to their failure. Such “mistakes” would more than likely produce a nonfunctional protein. Indeed, the cause of many genetic diseases is alternative splicing rather than mutations in a sequence. However, alternative splicing would create a protein variant without the loss of the original protein, opening up possibilities for adaptation of the new variant to new functions. Gene duplication has played an important role in the evolution of new functions in a similar way—by providing genes that may evolve without eliminating the original functional protein.
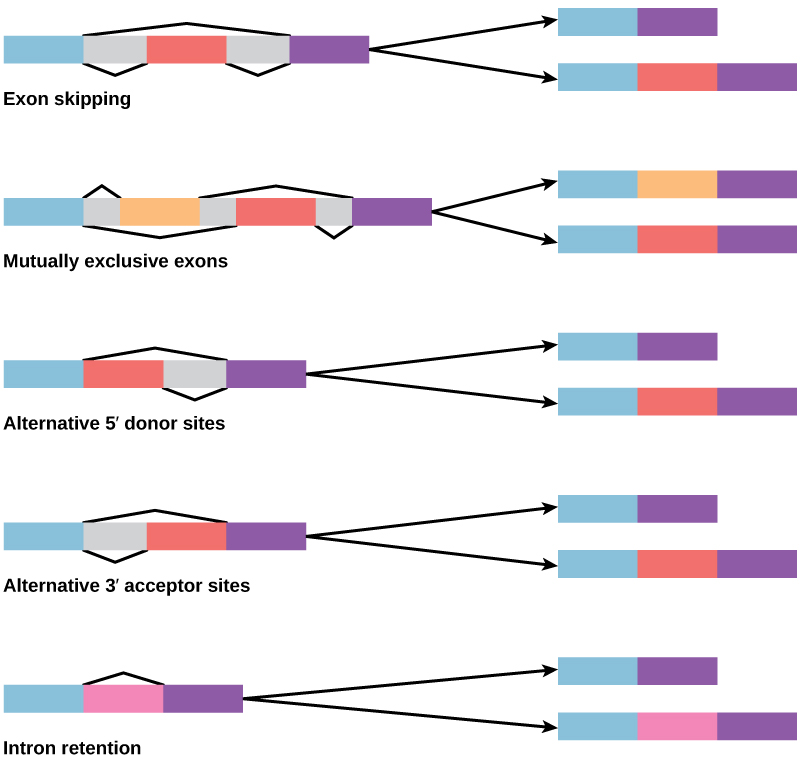
Control of RNA Stability
Before the mRNA leaves the nucleus, it is given two protective “caps” that prevent the end of the strand from degrading during its journey. The 5′ cap, which is placed on the 5′ end of the mRNA, is usually composed of a methylated guanosine triphosphate molecule (GTP). The poly-A tail, which is attached to the 3′ end, is usually composed of a series of adenine nucleotides. Once the RNA is transported to the cytoplasm, the length of time that the RNA remains there can be controlled. Each RNA molecule has a defined lifespan and decays at a specific rate. This rate of decay can influence how much protein is in the cell. If the RNA decays more rapidly, translation has less time to occur, so less protein will be produced. Conversely, if RNA decays less rapidly, more protein will be produced. This rate of decay is referred to as the RNA stability. If the RNA is stable, it will be detected for longer periods of time in the cytoplasm. Binding of proteins to the RNA can influence its stability (Figure 3).

Translational Regulation
Like transcription, translation is controlled by proteins that bind and initiate the process. In translation, the complex that assembles to start the process is referred to as the initiation complex. Regulation of the formation of this complex can increase or decrease rates of translation.
Post-translational Regulation
Proteins can be chemically modified with the addition of groups including methyl, phosphate, acetyl, and ubiquitin groups. The addition or removal of these groups from proteins regulates their activity or the length of time they exist in the cell. Sometimes these modifications can regulate where a protein is found in the cell—for example, in the nucleus, the cytoplasm, or attached to the plasma membrane.
Chemical modifications occur in response to external stimuli such as stress, the lack of nutrients, heat, or ultraviolet light exposure. These changes can alter epigenetic accessibility, transcription, mRNA stability, or translation—all resulting in changes in expression of various genes. This is an efficient way for the cell to rapidly change the levels of specific proteins in response to the environment. Because proteins are involved in every stage of gene regulation, the phosphorylation of a protein (depending on the protein that is modified) can alter accessibility to the chromosome, can alter translation (by altering transcription factor binding or function), can change nuclear shuttling (by influencing modifications to the nuclear pore complex), can alter RNA stability (by binding or not binding to the RNA to regulate its stability), can modify translation (increase or decrease), or can change post-translational modifications (add or remove phosphates or other chemical modifications).
The addition of an ubiquitin group to a protein marks that protein for degradation. Ubiquitin acts like a flag indicating that the protein lifespan is complete. These proteins are moved to the proteasome, an organelle that functions to remove proteins, to be degraded (Figure 2). One way to control gene expression, therefore, is to alter the longevity of the protein.

References
Unless otherwise noted, images on this page are licensed under CC-BY 4.0 by OpenStax.
OpenStax, Concepts of Biology. OpenStax CNX. January 3, 2017. https://cnx.org/contents/GFy_h8cu@10.120:7Ry3oRse@6
